All dwi data is available for downloads in the DWI page.
All tractograms are available for downloads in the Tools page.
Data generation in 2015
-
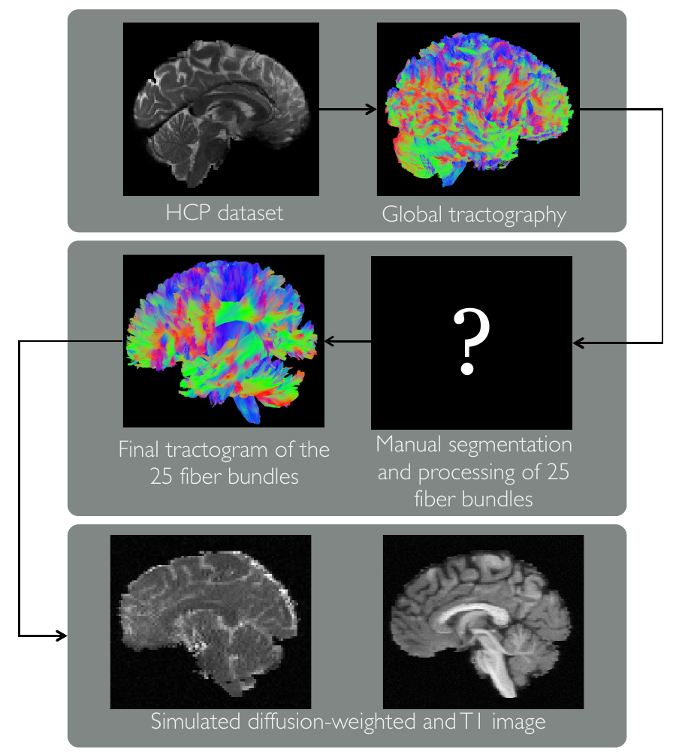
White matter bundles were manually segmentated from a HCP subject’s tractogram based on definitions found in Diffusion Tensor Imaging, Introduction and Atlas, which was written by challenge coorganizers Bram Stieltjes and Klaus Maier-Hein, as well as R.M. Brunner and F.B. Laun. To create these 25 bundles, an expert segmented meticulously streamlines of a global tracking on HCP data, the bundle masks were computed, and bundle-specific deterministic tracking was performed by seeding extensively inside each bundle mask.
-
The final tractogram was then used in Fiberfox to simulate the fitting DWI that can be found in the DWI data page, as described in the Fiberfox paper by Neher et al. The aim was to create a realistic, clinical-style dataset that provided challenging bundles configurations.
-
In 2023, to allow the creation of good Regions of Interest (ROIs), the ground truth tractogram was curated in Renauld2023 as it contained short / long / looping / broken streamlines. This data corresponds less with the simulated DWI but is also available for downloads for our most curious users.
The diffusion acquisition was simulated with the following parameters:
- Phase direction: Y
- No acceleration factor
- No partial Fourier
- TE = 108ms
- Dwell time = 1ms
** Important note about the Field Map: Please note that the same fieldmap is applied to all image volumes, regardless of head motion. This is a limitation of the current generation technique. In reality, the fieldmap should move with the head, but this is currently not the case in the simulated dataset.
The new filtered data is offered, together with the new scoring system, in the Tools page.